Current Issue
Display Method: |
Guest Editor:
Wenjun Mao The Affiliated Wuxi People’s Hospital of Nanjing Medical University, China
Xudong Qu Zhongshan Hospital, Fudan University, China
2026, 38(2): 119-120.
doi: 10.21147/j.issn.1000-9604.2026.02.01
Abstract:
2026, 38(2): 121-147.
doi: 10.21147/j.issn.1000-9604.2026.02.02
Abstract:
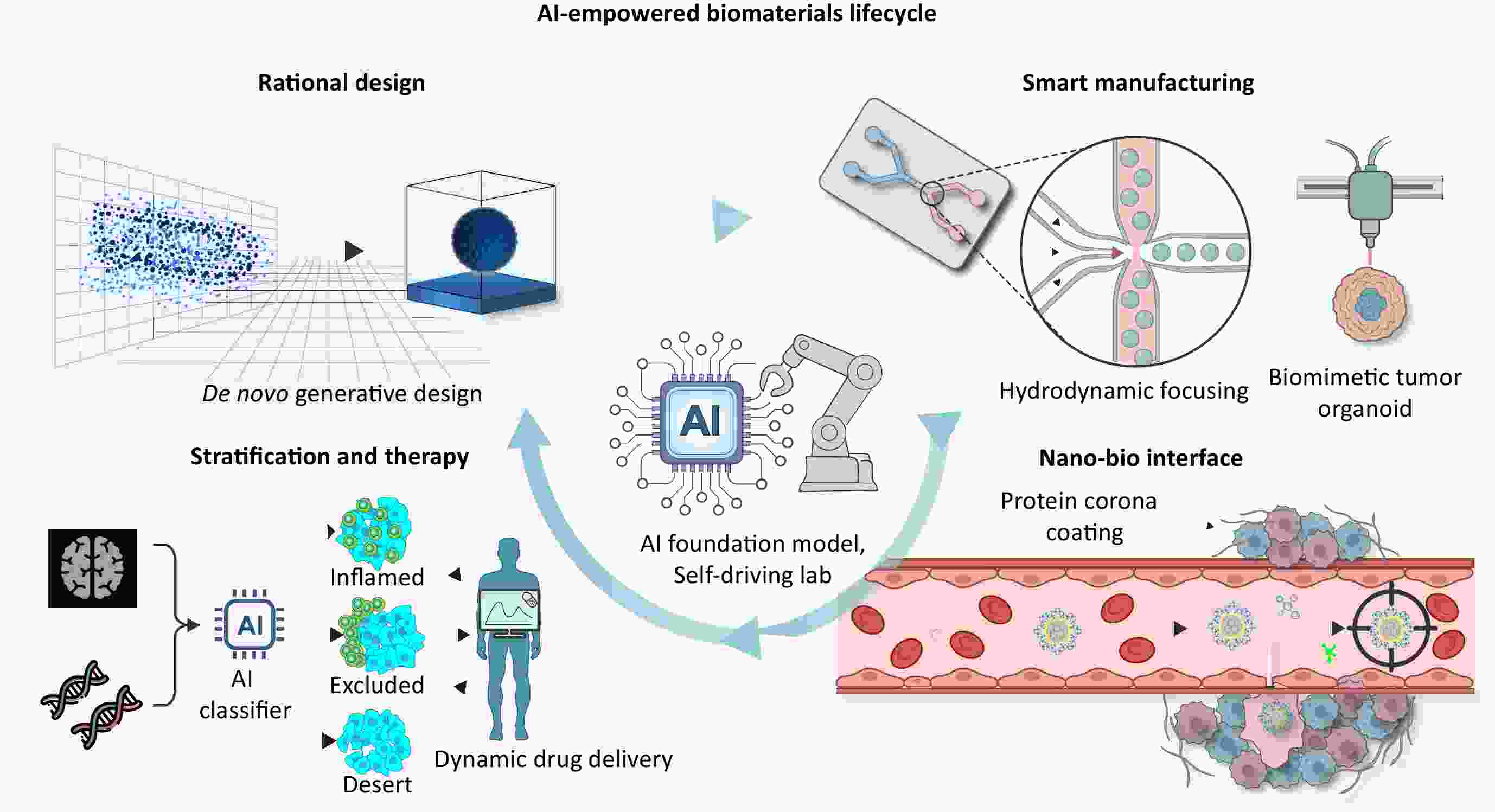
Tumor biomaterials show great potential for targeted cancer therapy, yet their development and clinical translation have long been hampered by inefficient empirical trial-and-error models. These traditional methods cannot fully characterize the nonlinear relationships between a material’s physicochemical properties and its complex biological effects, nor can they resolve tumor heterogeneity—the primary cause of inconsistent clinical outcomes. This review systematically explores the application of artificial intelligence (AI) across the entire development pipeline of tumor biomaterials, from early rational material design to clinical treatment optimization. We show that AI addresses key bottlenecks in the field in four core ways: it speeds up novel material discovery via generative algorithms, accurately predicts the in vivo transport and uptake of materials, enables noninvasive and precise patient stratification, and optimizes synergistic combination treatment regimens. These advances form a data-driven closed-loop framework that connects preclinical research and clinical translation, overcoming the core limitations of traditional development models. We also outline key unresolved challenges, including data standardization, model interpretability, and regulatory compliance, and highlight AI’s growing role as a core driver of precision oncology and translational medicine.
Tumor biomaterials show great potential for targeted cancer therapy, yet their development and clinical translation have long been hampered by inefficient empirical trial-and-error models. These traditional methods cannot fully characterize the nonlinear relationships between a material’s physicochemical properties and its complex biological effects, nor can they resolve tumor heterogeneity—the primary cause of inconsistent clinical outcomes. This review systematically explores the application of artificial intelligence (AI) across the entire development pipeline of tumor biomaterials, from early rational material design to clinical treatment optimization. We show that AI addresses key bottlenecks in the field in four core ways: it speeds up novel material discovery via generative algorithms, accurately predicts the in vivo transport and uptake of materials, enables noninvasive and precise patient stratification, and optimizes synergistic combination treatment regimens. These advances form a data-driven closed-loop framework that connects preclinical research and clinical translation, overcoming the core limitations of traditional development models. We also outline key unresolved challenges, including data standardization, model interpretability, and regulatory compliance, and highlight AI’s growing role as a core driver of precision oncology and translational medicine.
2026, 38(2): 148-176.
doi: 10.21147/j.issn.1000-9604.2026.02.03
Abstract:
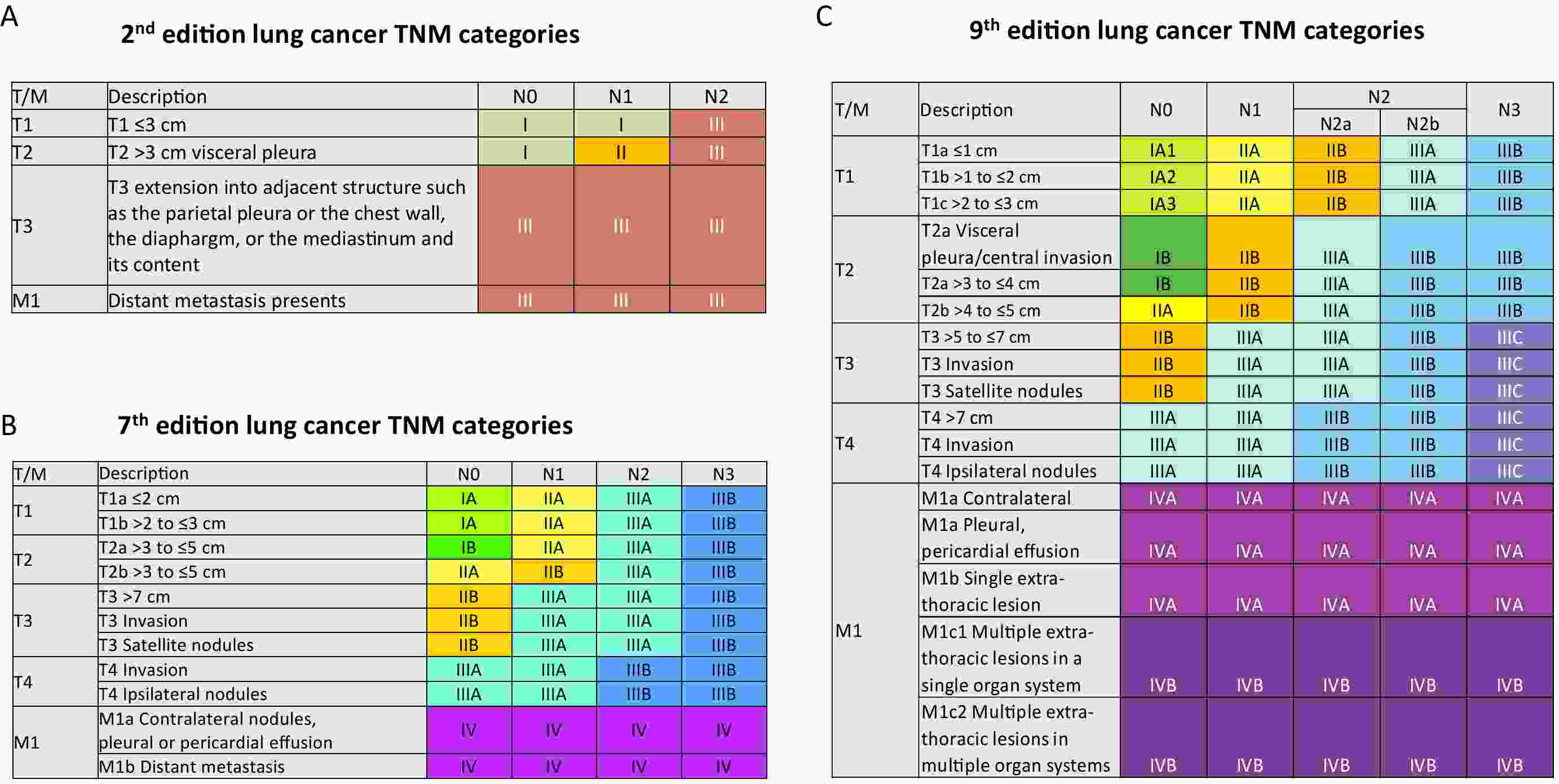
Lung cancer is the most lethal malignancy worldwide, largely due to its late detection after its progression to advanced stages. Over the last decade, artificial intelligence (AI) applications have shown significant potential in transforming lung cancer diagnostics by improving the speed, accuracy, and personalization of early detection strategies. This review provides a comprehensive overview of current AI application landscape in early lung cancer diagnosis, encompassing medical imaging, histopathology, liquid biopsy, natural language processing of electronic health records, and genomic profiling. We explain how machine learning, deep learning, and transformer-based models are employed in lung cancer diagnosis, and summarize recent cutting-edge advances, including multimodal AI platforms and Food and Drug Administration (FDA)-approved computer-aided diagnosis/detection (CAD) systems. Furthermore, we evaluate the challenges that impede clinical translation, including data heterogeneity, interpretability, and privacy, and present prospective directions such as federated learning and multi-omics integration. Through a comprehensive analysis of the dynamic evolution of AI applications in oncology, we aim to inform researchers, clinicians, and policymakers about its diagnostic potential and translational relevance in clinical practice.
Lung cancer is the most lethal malignancy worldwide, largely due to its late detection after its progression to advanced stages. Over the last decade, artificial intelligence (AI) applications have shown significant potential in transforming lung cancer diagnostics by improving the speed, accuracy, and personalization of early detection strategies. This review provides a comprehensive overview of current AI application landscape in early lung cancer diagnosis, encompassing medical imaging, histopathology, liquid biopsy, natural language processing of electronic health records, and genomic profiling. We explain how machine learning, deep learning, and transformer-based models are employed in lung cancer diagnosis, and summarize recent cutting-edge advances, including multimodal AI platforms and Food and Drug Administration (FDA)-approved computer-aided diagnosis/detection (CAD) systems. Furthermore, we evaluate the challenges that impede clinical translation, including data heterogeneity, interpretability, and privacy, and present prospective directions such as federated learning and multi-omics integration. Through a comprehensive analysis of the dynamic evolution of AI applications in oncology, we aim to inform researchers, clinicians, and policymakers about its diagnostic potential and translational relevance in clinical practice.
2026, 38(2): 177-196.
doi: 10.21147/j.issn.1000-9604.2026.02.04
Abstract:
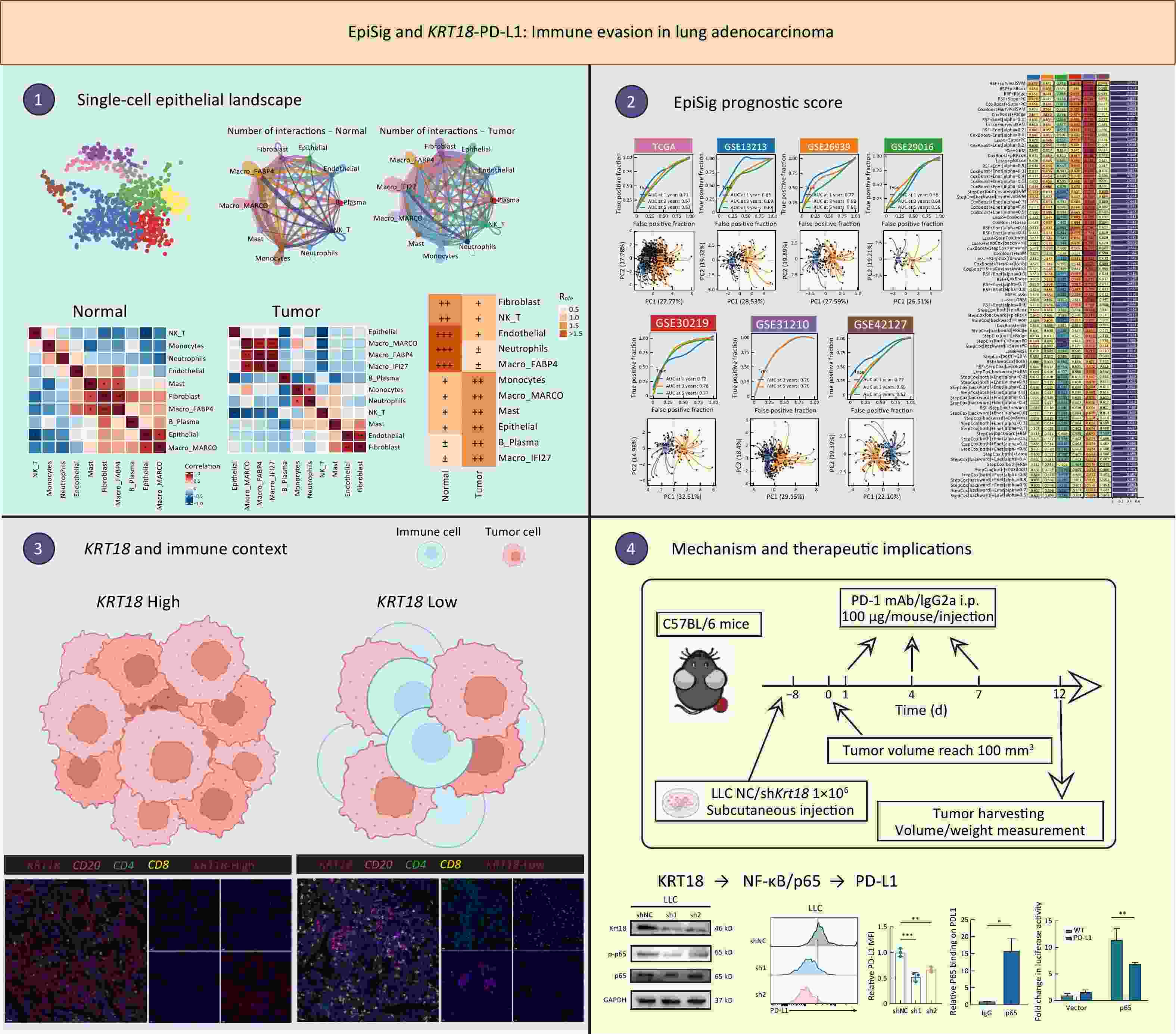
ObjectiveThe heterogeneity of epithelial cells and their interaction with the immune microenvironment play crucial roles in tumor progression, but the underlying mechanisms remain unclear. MethodsWe analyzed single-cell transcriptomic data from normal and tumor tissues to characterize epithelial cells and their microenvironment. Key genes were identified and used, via survival analysis and multiple machine learning methods, to construct a prognostic model termed the Epithelial Signature (EpiSig). We further validated, through a series of experiments, the critical immunological roles of the key genes incorporated into the EpiSig model. ResultsTumor tissues showed a marked increase in epithelial cells, a reduction in natural killer (NK)/T cells, and cell co-occurrence patterns distinct from normal tissues. We identified differentially expressed genes in tumor epithelial cells and integrated multiple machine-learning algorithms to construct the EpiSig model. This model effectively stratified patient prognosis, with the high-EpiSig group exhibiting significantly worse survival; receiver operator characteristic curve (ROC) and principal component analysis (PCA) analyses further supported its accuracy and robustness. Immune analyses indicated lower immune cell infiltration, decreased human leukocyte antigen (HLA) expression, and elevated programmed cell death ligand 1/programmed cell death protein 1 (PD-L1/PD-1) in the high-EpiSig group, reflecting a more pronounced immunosuppressive microenvironment. The core gene KRT18 correlated strongly with EpiSig scores (R=0.65) and promoted lung adenocarcinoma (LUAD) cell proliferation in functional assays. Further, low KRT18 tumors showed higher CD4+/CD8+ T cell and CD20+ B cell infiltration; in a mouse LLC subcutaneous tumor model, both Krt18 knockdown and PD-1 blockade suppressed tumor growth, with greater efficacy in combination. Mechanistically, KRT18 activated NF-kB/p65 to upregulate PD-L1, promoting immune exclusion and impairing T-cell effector function. ConclusionsThis study highlights the close relationship between epithelial cell heterogeneity and immune microenvironment alterations in tumors, and presents the EpiSig as a robust tool for prognostic prediction. KRT18 may serve as a promising therapeutic target in LUAD.
2026, 38(2): 209-233.
doi: 10.21147/j.issn.1000-9604.2026.02.08
Abstract:
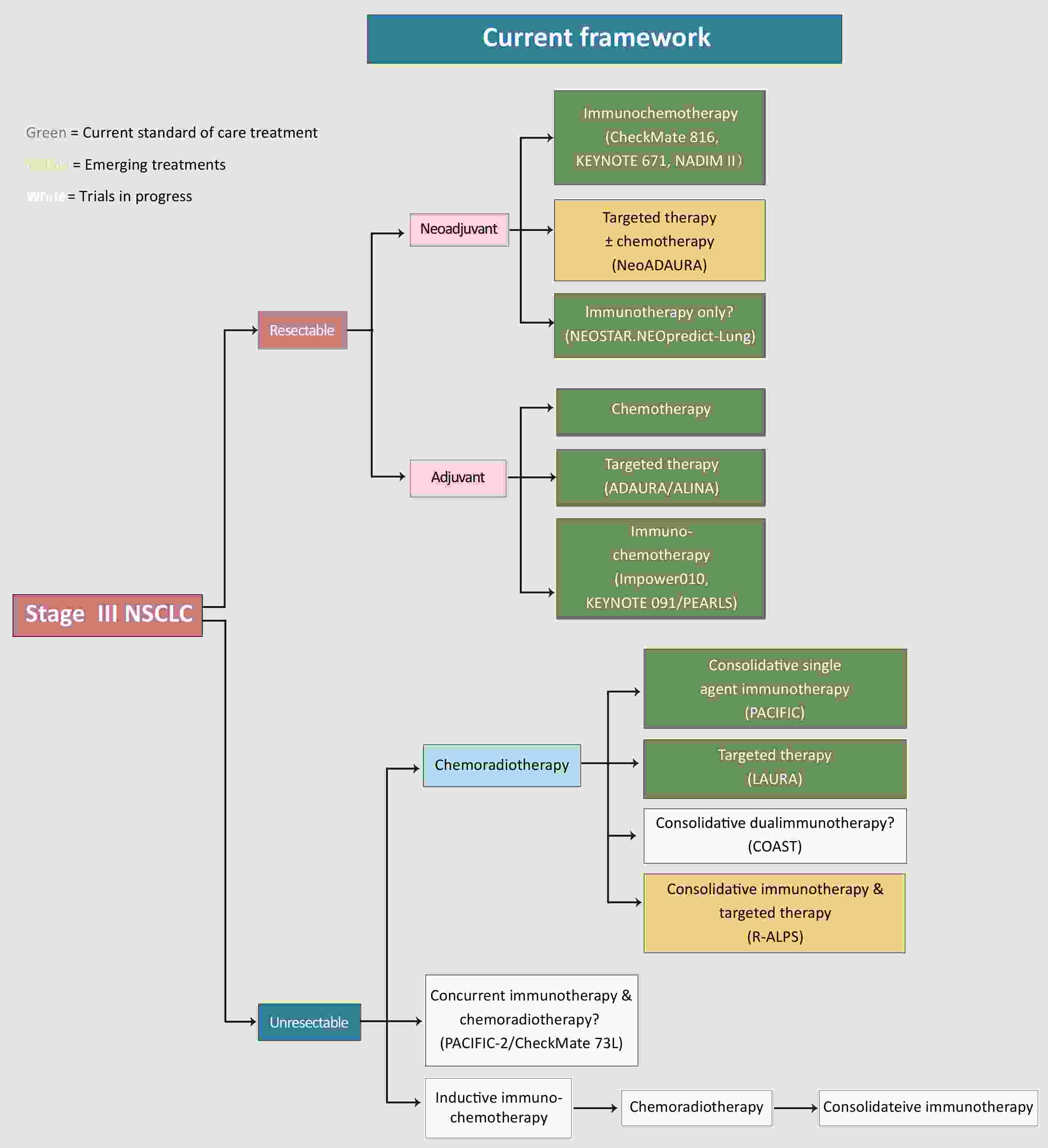
Stage III non-small cell lung cancer (NSCLC) is traditionally viewed as a locally advanced disease based on anatomic imaging. It is increasingly recognized as a systemic illness with localized manifestations, as evidenced by high rates of distant recurrence despite aggressive local therapy. Recent landmark trials (e.g., ADAURA, ALINA, PACIFIC, and KEYNOTE 671) have demonstrated that prolonged systemic treatment, including adjuvant targeted therapy in oncogene-driven NSCLC and consolidation immune checkpoint inhibition after chemoradiotherapy, significantly improves survival outcomes. In contrast, intensifying local therapy alone has exhibited limited benefit or even potential harm. Supporting this shift, detection of circulating tumor DNA and circulating tumor cells indicates the presence of micrometastatic disease at the time of diagnosis. We propose an integrated “sandwich” therapeutic framework encompassing three sequential phases: 1) induction with biomarker-guided systemic therapy (e.g., targeted agents or chemoimmunotherapy) to control micrometastases; 2) local consolidation with surgery or radiotherapy tailored to the post-induction tumor extent; and 3) systemic consolidation with prolonged maintenance therapy to eradicate residual disease. This approach underscores the necessity of treating stage III NSCLC as a systemic disease from the outset, integrating prolonged, biomarker-directed systemic strategies within a multimodal curative-intent framework to address both the local and systemic components of the disease.
Stage III non-small cell lung cancer (NSCLC) is traditionally viewed as a locally advanced disease based on anatomic imaging. It is increasingly recognized as a systemic illness with localized manifestations, as evidenced by high rates of distant recurrence despite aggressive local therapy. Recent landmark trials (e.g., ADAURA, ALINA, PACIFIC, and KEYNOTE 671) have demonstrated that prolonged systemic treatment, including adjuvant targeted therapy in oncogene-driven NSCLC and consolidation immune checkpoint inhibition after chemoradiotherapy, significantly improves survival outcomes. In contrast, intensifying local therapy alone has exhibited limited benefit or even potential harm. Supporting this shift, detection of circulating tumor DNA and circulating tumor cells indicates the presence of micrometastatic disease at the time of diagnosis. We propose an integrated “sandwich” therapeutic framework encompassing three sequential phases: 1) induction with biomarker-guided systemic therapy (e.g., targeted agents or chemoimmunotherapy) to control micrometastases; 2) local consolidation with surgery or radiotherapy tailored to the post-induction tumor extent; and 3) systemic consolidation with prolonged maintenance therapy to eradicate residual disease. This approach underscores the necessity of treating stage III NSCLC as a systemic disease from the outset, integrating prolonged, biomarker-directed systemic strategies within a multimodal curative-intent framework to address both the local and systemic components of the disease.
2026, 38(2): 234-251.
doi: 10.21147/j.issn.1000-9604.2026.02.09
Abstract:
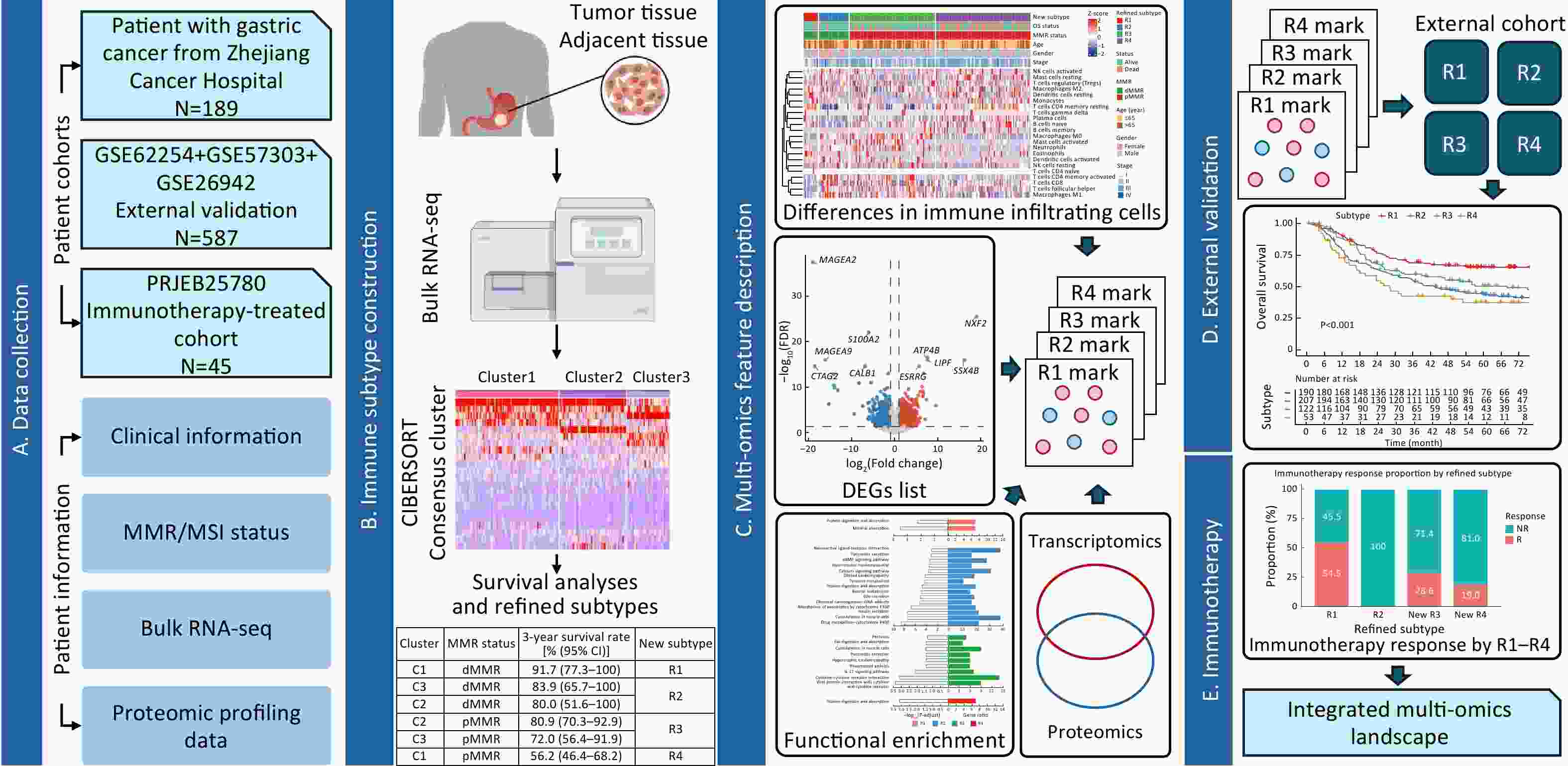
ObjectiveGastric cancer (GC) is heterogeneous, and current mismatch repair (MMR)-based classifications incompletely predict response to immune checkpoint inhibitors (ICIs). MethodsRNA sequencing (RNA-seq) and immune infiltration profiles from 189 resected GC were used to derive four refined immune-MMR subtypes (R1−R4) by integrating MMR status, survival, and tumor microenvironment (TME) features. Multi-omics profiling and pathway analysis defined subtype biology. External transcriptomic cohorts and an ICI-treated cohort were classified with Nearest Template Prediction (NTP). Immune response-associated genes were identified from responder vs. non-responder comparisons within the ICI-sensitive subtype and validated by multiplex immunohistochemistry (mIHC). ResultsR1 showed the best prognosis and highest immunotherapy response with objective response rate (ORR) 54.5%, while R4 had the worst prognosis. R2 represented an immune-unresponsive deficient mismatch repair (dMMR) subset, and R3 captured an immune-active proficient mismatch repair (pMMR) subgroup with moderate therapy sensitivity. Multi-omics integration revealed subtype-specific pathways (e.g., ECM remodeling in R1, metabolic reprogramming in R2). Reclassification of pMMR tumors based on transcriptional similarity to R1 identified a New R3 subset with enhanced immune features and higher ICI response. Eight immune response-associated genes (e.g., CXCL10, CXCL11, ELN, GAD1, IL32, MT1E, OR2I1P, SLC3A1) were identified and validated by mIHC for predictive relevance. ConclusionsThis immune-based molecular framework refines risk stratification beyond conventional MMR categories, identifies ICI-sensitive subsets among both dMMR and pMMR tumors, and proposes candidate biomarkers for patient selection.
2026, 38(2): 252-267.
doi: 10.21147/j.issn.1000-9604.2026.02.10
Abstract:
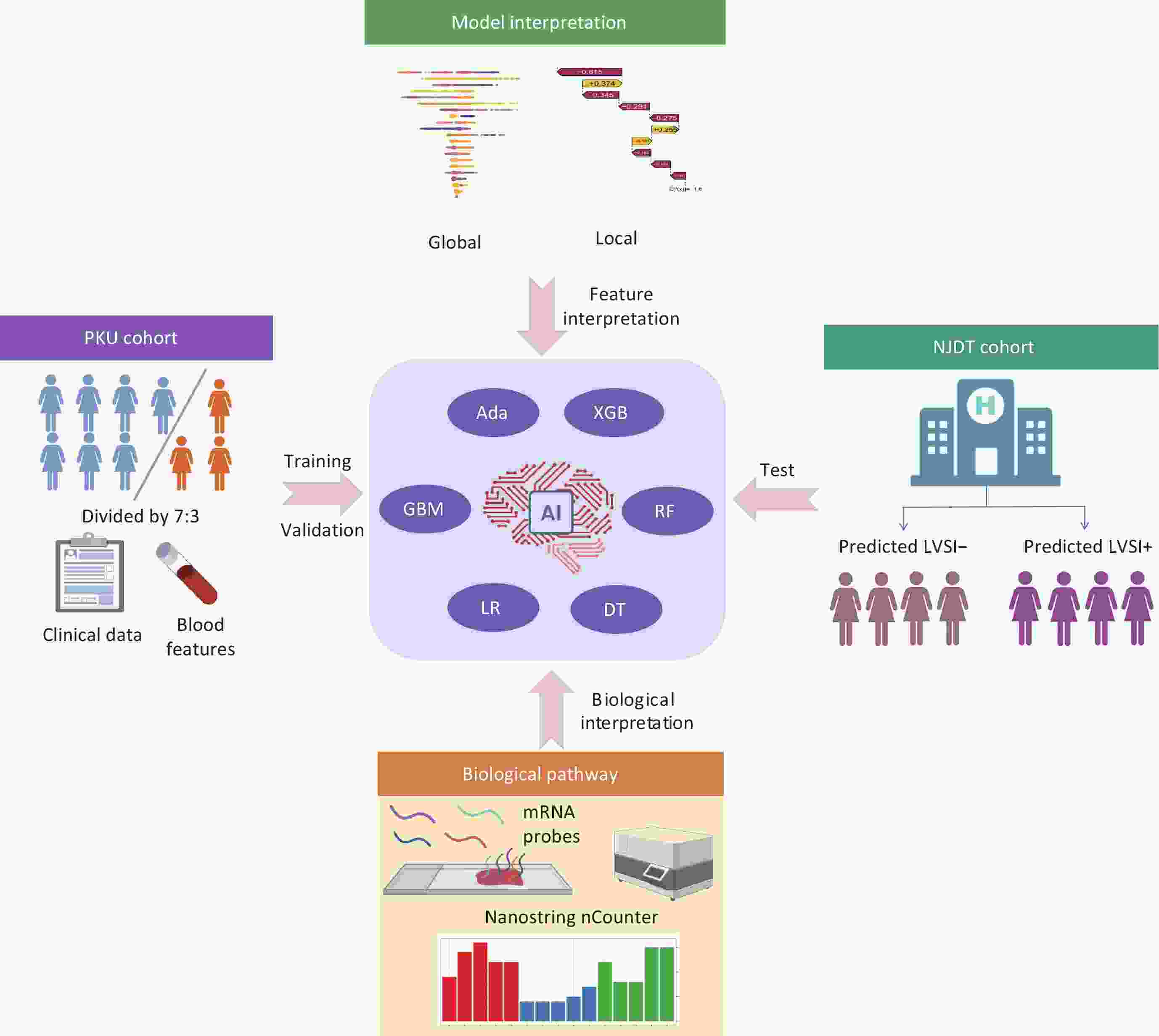
ObjectiveLymphovascular space invasion (LVSI) is a high-risk factor for lymph node metastasis, relapse, and poor prognosis in patients with endometrioid endometrial carcinoma (EEC). However, the diagnosis of LVSI still relies on traditional pathological methods. Moreover, the high-risk factors and mechanism for LVSI remain unclear. Thus, this study developed an interpretable machine learning (ML) model to accurately predict LVSI status in patients with EEC. MethodsThe study collected data from 832 patients with EEC at Peking University People’s Hospital. Patients were randomly divided into training (n=582) and internal validation (n=250) cohorts. A prospective external validation cohort included 129 patients with EEC from Nanjing Drum Tower Hospital. Using 21 parameters, 6 ML strategies were used to build prediction models. The global and local interpretation of feature significance was performed using the SHapley Additive exPlanations (SHAP) approach. Data from NanoString nCounter evaluation was subjected to pathway enrichment and Spearman correlation analysis to investigate the mechanistic basis of LVSI. ResultsAmong the six ML models, the XGBoost model had the best performance. The XGBoost model correctly predicted the risk of LVSI in the training set [area under the curve (AUC): 0.982, 95% confidence interval (95% CI): 0.972−0.991], the internal validation set (AUC: 0.818, 95% CI: 0.776−0.860), and the external test set (AUC: 0.748, 95% CI: 0.618−0.879). The calibration curve indicated that the XGBoost model exhibited favorable consistency between the predicted and actual risks. SHAP analysis identified age, carbohydrate antigen 125 (CA125), low-density lipoprotein (LDL), and neutrophil as the top four variables contributing to XGBoost model predictions. Analysis of NanoString data indicated that LVSI may be closely associated with the PI3K-Akt signaling pathway. ConclusionsWe developed an interpretable ML model for preoperative LVSI risk prediction in patients with EEC. This model may aid clinicians by informing individualized clinical decision-making.
2026, 38(2): 268-284.
doi: 10.21147/j.issn.1000-9604.2026.02.11
Abstract:
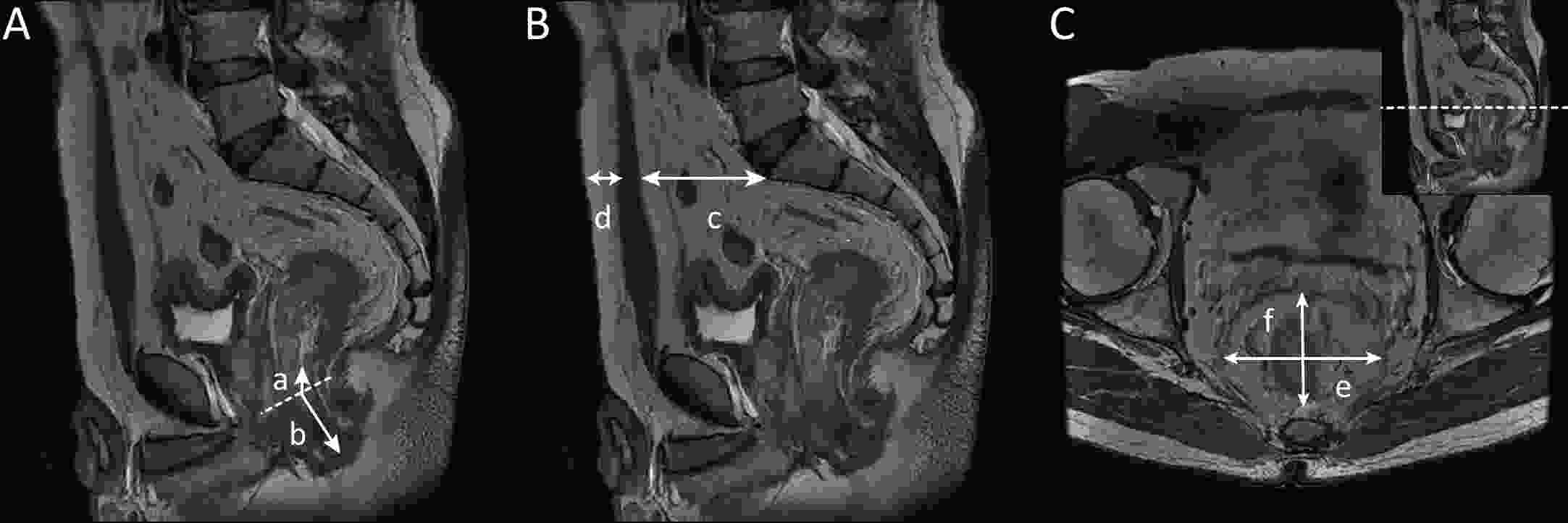
ObjectiveAlthough distance from the inferior tumor edge to the anal verge (DTAV) is a key predictor for sphincter-preserving surgery (SPS) in mid-low rectal cancer, its utility is limited in the “decision-gray zone” (DTAV, 3−8 cm). Therefore, this study aimed to develop and validate a multiparametric magnetic resonance imaging-based nomogram for individualized preoperative prediction of SPS feasibility. MethodsThis dual-center retrospective study included 335 patients with rectal adenocarcinoma (DTAV 3−8 cm). Patients were divided into training (n=263) and external validation (n=72) cohorts, and predictors were identified using multivariate logistic regression analysis. Model discrimination was assessed using area under the receiver operating curve (AUC) and calibration via the Hosmer-Lemeshow test. Subgroup analyses were performed across DTAV strata. ResultsFour independent predictors were identified: larger DTAV [odds ratio (OR) =5.00, P<0.001)], larger pubococcygeal overlap distance (PCOD) (OR=1.08, P=0.001), transverse diameter of mesorectal fat (TMS) (OR=1.07, P=0.017), and subcutaneous adipose tissue thickness (SAT) (OR=0.94, P=0.016). The Sphincter Preservation Assessment in Rectal Cancer (SPARC) nomogram achieved an AUC of 0.928 [95% confidence interval (95% CI): 0.890−0.956] in the training cohort, outperforming DTAV alone (AUC=0.884, P=0.031) and maintaining an AUC of 0.916 (95% CI: 0.827−0.969) in external validation. Subgroup analysis showed notably improved predictions in the 5−8 cm DTAV subgroup. Decision curve analysis demonstrated a pronounced net clinical benefit across a wide range of threshold probabilities. Interobserver agreement was excellent (intraclass correlation coefficient, 0.890−0.997). ConclusionsThe SPARC nomogram reliably predicted SPS feasibility by integrating tumor location with pelvic anatomy and fat distribution. This provides valuable and evidence-based preoperative guidance, especially within the DTAV 3−8 cm gray zone.
 Abstract
Abstract FullText HTML
FullText HTML PDF 824KB
PDF 824KB